
Anchoring: Covalently linking your imaging probe to the expansion gel itself such that can be embedded in the cross-linked network (Figure 3b) Intermediate steps within labeling include washing out unbound antibodies and ‘ blocking’ the sample with protein to minimize nonspecific antibody binding.Ģ. This way you can visualize the spatial distribution of that protein. The secondary antibody is conjugated to a fluorophore which can be probed during microscopy. 
Next, a secondary antibody that is specific to the primary is added to the sample.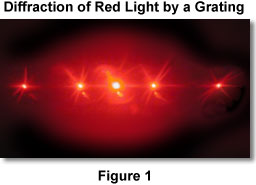 
Initially, the primary antibody is used to bind to your protein of interest. In immunostaining, antibodies are iteratively used to target proteins of interest. Labeling is typically performed using general immunostaining methods. Labeling: Attaching an imaging probe to your proteins of interest for eventual visualization during imaging (Figure 3a) What are these steps and how do they work? Figure 2: Polyelectrolyte hydrogels' polymer network expands in the presence of waterġ. The general strategy of ExM, developed by Ed Boyden’s group at MIT, is to apply this principle to biological tissues through a series of chemical infusions and treatments which alter the properties of the tissue while preserving the cellular and protein structure. At the molecular level, this is caused by the isotropic expansion of the polymer network (Figure 2). One property of these gels is that under certain environmental conditions, like being placed in water, they can swell. Polyelectrolyte gels are composed of crosslinked polymers that form a grid network. How can AI integrate into the workflow of high-resolution imaging afforded by Expansion Microscopy?Įxpansion Microscopy deploys hydrogel chemistry to isotropically expand biological tissues How is Expansion Microscopy being used in fields like neuroscience and pathology?ģ. What is Expansion Microscopy and how does it work?Ģ. This determines how close two light sources can be disambiguated.ġ. Figure 1: Schematic representation of the diffraction limit of an optical lens (a) the numerical aperture (NA) of a lens is a function of the maximal angle that the lens can receive light (b) the diffraction limit is the fraction of the wavelength of the light and the numerical aperture. In this article, I will discuss how it works and how scientists are using it to cheat the diffraction limit and peer into previously inaccessible nanoscopic worlds. What if instead, we could literally grow the sample itself like consuming a super mushroom in Mario Bros? Scientists have developed a method whereby this is possible, and it is called Expansion Microscopy (ExM). High-resolution microscopes, however, are already incredibly close to the sample. In our thought experiment, the subject grew in the frame by becoming closer to the sensor. The resolution of this system, however, is limited by its diffraction limit which is a result of how light bends due to its wave properties. Mathematically, the resolution is a function of the numerical aperture of the optical system and the wavelength of the light you are collecting (figure 1). A paramount goal in the field of microscopy is to image increasingly smaller things at increasing degrees of resolution. This provides a useful framework for conceptualizing resolution, which measures how many pixels there are and how much information each pixel encodes. What did you just do? You decreased the distance between the camera sensor and the subject, such that now each pixel occupies less surface area and thus represents finer information. To adjust for this, you move in closer and snap the perfect picture. Where you stand, you notice that although you are capturing much of the background, your friend is far away and consequently they appear blurry. Imagine you are behind the camera framing a picture of your friend.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
